Simple Introduction
This tutorial demonstrates the capability to perform ensembles of calculations in parallel using libEnsemble.
We recommend reading this brief Overview.
For this tutorial, our generator will produce uniform randomly sampled values, and our simulator will calculate the sine of each. By default we don’t need to write a new allocation function.
libEnsemble is written entirely in Python. Let’s make sure the correct version is installed.
python --version # This should be >= 3.9
For this tutorial, you need NumPy and (optionally) Matplotlib to visualize your results. Install libEnsemble and these other libraries with
pip install libensemble
pip install matplotlib # Optional
If your system doesn’t allow you to perform these installations, try adding
--user to the end of each command.
Let’s begin the coding portion of this tutorial by writing our generator function, or gen_f.
An available libEnsemble worker will call this generator function with the following parameters:
InputArray: A selection of the History array (H), passed to the generator function in case the user wants to generate new values based on simulation outputs. Since our generator produces random numbers, it’ll be ignored this time.
persis_info: Dictionary with worker-specific information. In our case, this dictionary contains NumPy Random Stream objects for generating random numbers.
gen_specs: Dictionary with user-defined static fields and parameters. Customizable parameters such as lower and upper bounds and batch sizes are placed within the
gen_specs["user"]dictionary.
Later on, we’ll populate gen_specs and persis_info when we initialize libEnsemble.
For now, create a new Python file named sine_gen.py. Write the following:
1import numpy as np
2
3
4def gen_random_sample(InputArray, persis_info, gen_specs):
5 # Pull out user parameters
6 user_specs = gen_specs["user"]
7
8 # Get lower and upper bounds
9 lower = user_specs["lower"]
10 upper = user_specs["upper"]
11
12 # Determine how many values to generate
13 num = len(lower)
14 batch_size = user_specs["gen_batch_size"]
15
16 # Create empty array of "batch_size" zeros. Array dtype should match "out" fields
17 OutputArray = np.zeros(batch_size, dtype=gen_specs["out"])
18
19 # Set the "x" output field to contain random numbers, using random stream
20 OutputArray["x"] = persis_info["rand_stream"].uniform(lower, upper, (batch_size, num))
21
22 # Send back our output and persis_info
23 return OutputArray, persis_info
Our function creates batch_size random numbers uniformly distributed
between the lower and upper bounds. A random stream
from persis_info is used to generate these values, which are then placed
into an output NumPy array that matches the dtype from gen_specs["out"].
Next, we’ll write our simulator function or sim_f. Simulator
functions perform calculations based on values from the generator function.
The only new parameter here is sim_specs, which
serves a purpose similar to the gen_specs dictionary.
Create a new Python file named sine_sim.py. Write the following:
1import numpy as np
2
3
4def sim_find_sine(InputArray, _, sim_specs):
5 # Create an output array of a single zero
6 OutputArray = np.zeros(1, dtype=sim_specs["out"])
7
8 # Set the zero to the sine of the InputArray value
9 OutputArray["y"] = np.sin(InputArray["x"])
10
11 # Send back our output
12 return OutputArray
Our simulator function is called by a worker for every work item produced by the generator function. This function calculates the sine of the passed value, and then returns it so the worker can store the result.
Now lets write the script that configures our generator and simulator functions and starts libEnsemble.
Create an empty Python file named calling.py.
In this file, we’ll start by importing NumPy, libEnsemble’s setup classes,
and the generator and simulator functions we just created.
In a class called LibeSpecs we’ll
specify the number of workers and the manager/worker intercommunication method.
"local", refers to Python’s multiprocessing.
1import numpy as np
2from sine_gen import gen_random_sample
3from sine_sim import sim_find_sine
4
5from libensemble import Ensemble
6from libensemble.specs import ExitCriteria, GenSpecs, LibeSpecs, SimSpecs
7
8if __name__ == "__main__": # Python-quirk required on macOS and windows
9 libE_specs = LibeSpecs(nworkers=4, comms="local")
We configure the settings and specifications for our sim_f and gen_f
functions in the GenSpecs and
SimSpecs classes, which we saw previously
being passed to our functions as dictionaries.
These classes also describe to libEnsemble what inputs and outputs from those
functions to expect.
10 gen_specs = GenSpecs(
11 gen_f=gen_random_sample, # Our generator function
12 out=[("x", float, (1,))], # gen_f output (name, type, size)
13 user={
14 "lower": np.array([-3]), # lower boundary for random sampling
15 "upper": np.array([3]), # upper boundary for random sampling
16 "gen_batch_size": 5, # number of x's gen_f generates per call
17 },
18 )
19
20 sim_specs = SimSpecs(
21 sim_f=sim_find_sine, # Our simulator function
22 inputs=["x"], # InputArray field names. "x" from gen_f output
23 out=[("y", float)], # sim_f output. "y" = sine("x")
24 ) # sim_specs_end_tag
We then specify the circumstances where libEnsemble should stop execution in ExitCriteria.
26 exit_criteria = ExitCriteria(sim_max=80) # Stop libEnsemble after 80 simulations
Now we’re ready to write our libEnsemble libE
function call. ensemble.H is the final version of
the history array. ensemble.flag should be zero if no errors occur.
28 ensemble = Ensemble(sim_specs, gen_specs, exit_criteria, libE_specs)
29 ensemble.add_random_streams() # setup the random streams unique to each worker
30 ensemble.run() # start the ensemble. Blocks until completion.
31
32 history = ensemble.H # start visualizing our results
33
34 print([i for i in history.dtype.fields]) # (optional) to visualize our history array
35 print(history)
That’s it! Now that these files are complete, we can run our simulation.
python calling.py
If everything ran perfectly and you included the above print statements, you should get something similar to the following output (although the columns might be rearranged).
["y", "sim_started_time", "gen_worker", "sim_worker", "sim_started", "sim_ended", "x", "allocated", "sim_id", "gen_ended_time"]
[(-0.37466051, 1.559+09, 2, 2, True, True, [-0.38403059], True, 0, 1.559+09)
(-0.29279634, 1.559+09, 2, 3, True, True, [-2.84444261], True, 1, 1.559+09)
( 0.29358492, 1.559+09, 2, 4, True, True, [ 0.29797487], True, 2, 1.559+09)
(-0.3783986, 1.559+09, 2, 1, True, True, [-0.38806564], True, 3, 1.559+09)
(-0.45982062, 1.559+09, 2, 2, True, True, [-0.47779319], True, 4, 1.559+09)
...
In this arrangement, our output values are listed on the far left with the generated values being the fourth column from the right.
Two additional log files should also have been created.
ensemble.log contains debugging or informational logging output from
libEnsemble, while libE_stats.txt contains a quick summary of all
calculations performed.
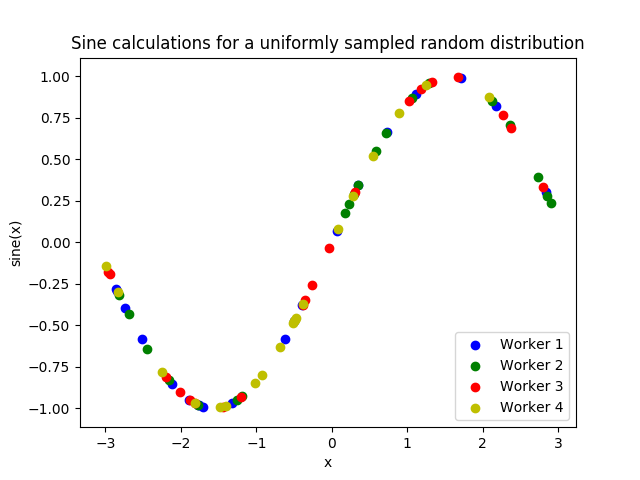
Here is graphed output using Matplotlib, with entries colored by which
worker performed the simulation:
If you want to verify your results through plotting and installed Matplotlib
earlier, copy and paste the following code into the bottom of your calling
script and run python calling.py again
37 import matplotlib.pyplot as plt
38
39 colors = ["b", "g", "r", "y", "m", "c", "k", "w"]
40
41 for i in range(1, libE_specs.nworkers + 1):
42 worker_xy = np.extract(history["sim_worker"] == i, history)
43 x = [entry.tolist()[0] for entry in worker_xy["x"]]
44 y = [entry for entry in worker_xy["y"]]
45 plt.scatter(x, y, label="Worker {}".format(i), c=colors[i - 1])
46
47 plt.title("Sine calculations for a uniformly sampled random distribution")
48 plt.xlabel("x")
49 plt.ylabel("sine(x)")
50 plt.legend(loc="lower right")
51 plt.savefig("tutorial_sines.png")
Each of these example files can be found in the repository in examples/tutorials/simple_sine.
Exercise
Write a Calling Script with the following specifications:
Set the generator function’s lower and upper bounds to -6 and 6, respectively
Increase the generator batch size to 10
Set libEnsemble to stop execution after 160 generations using the
gen_maxoptionPrint an error message if any errors occurred while libEnsemble was running
Click Here for Solution
1import numpy as np
2from sine_gen import gen_random_sample
3from sine_sim import sim_find_sine
4
5from libensemble import Ensemble
6from libensemble.specs import ExitCriteria, GenSpecs, LibeSpecs, SimSpecs
7
8if __name__ == "__main__":
9 libE_specs = LibeSpecs(nworkers=4, comms="local")
10
11 gen_specs = GenSpecs(
12 gen_f=gen_random_sample, # Our generator function
13 out=[("x", float, (1,))], # gen_f output (name, type, size)
14 user={
15 "lower": np.array([-6]), # lower boundary for random sampling
16 "upper": np.array([6]), # upper boundary for random sampling
17 "gen_batch_size": 10, # number of x's gen_f generates per call
18 },
19 )
20
21 sim_specs = SimSpecs(
22 sim_f=sim_find_sine, # Our simulator function
23 inputs=["x"], # InputArray field names. "x" from gen_f output
24 out=[("y", float)], # sim_f output. "y" = sine("x")
25 )
26
27 exit_criteria = ExitCriteria(gen_max=160)
28
29 ensemble = Ensemble(sim_specs, gen_specs, exit_criteria, libE_specs)
30 ensemble.add_random_streams()
31 ensemble.run()
32
33 if ensemble.flag != 0:
34 print("Oh no! An error occurred!")
libEnsemble with MPI
MPI is a standard interface for parallel computing, implemented in libraries such as MPICH and used at extreme scales. MPI potentially allows libEnsemble’s processes to be distributed over multiple nodes and works in some circumstances where Python’s multiprocessing does not. In this section, we’ll explore modifying the above code to use MPI instead of multiprocessing.
We recommend the MPI distribution MPICH for this tutorial, which can be found
for a variety of systems here. You also need mpi4py, which can be installed
with pip install mpi4py. If you’d like to use a specific version or
distribution of MPI instead of MPICH, configure mpi4py with that MPI at
installation with MPICC=<path/to/MPI_C_compiler> pip install mpi4py If this
doesn’t work, try appending --user to the end of the command. See the
mpi4py docs for more information.
Verify that MPI has been installed correctly with mpirun --version.
Modifying the script
Only a few changes are necessary to make our code MPI-compatible. For starters,
comment out the libE_specs definition:
# libE_specs = LibeSpecs(nworkers=4, comms="local")
We’ll be parameterizing our MPI runtime with a parse_args=True argument to
the Ensemble class instead of libE_specs. We’ll also use an ensemble.is_manager
attribute so only the first MPI rank runs the data-processing code.
The bottom of your calling script should now resemble:
28 # replace libE_specs with parse_args=True. Detects MPI runtime
29 ensemble = Ensemble(sim_specs, gen_specs, exit_criteria, parse_args=True)
30
31 ensemble.add_random_streams()
32 ensemble.run() # start the ensemble. Blocks until completion.
33
34 if ensemble.is_manager: # only True on rank 0
35 history = ensemble.H # start visualizing our results
36 print([i for i in history.dtype.fields])
37 print(history)
38
39 import matplotlib.pyplot as plt
40
41 colors = ["b", "g", "r", "y", "m", "c", "k", "w"]
42
43 for i in range(1, ensemble.nworkers + 1):
44 worker_xy = np.extract(history["sim_worker"] == i, history)
45 x = [entry.tolist()[0] for entry in worker_xy["x"]]
46 y = [entry for entry in worker_xy["y"]]
47 plt.scatter(x, y, label="Worker {}".format(i), c=colors[i - 1])
48
49 plt.title("Sine calculations for a uniformly sampled random distribution")
50 plt.xlabel("x")
51 plt.ylabel("sine(x)")
52 plt.legend(loc="lower right")
53 plt.savefig("tutorial_sines.png")
With these changes in place, our libEnsemble code can be run with MPI by
mpirun -n 5 python calling.py
where -n 5 tells mpirun to produce five processes, one of which will be
the manager process with the libEnsemble manager and the other four will run
libEnsemble workers.
This tutorial is only a tiny demonstration of the parallelism capabilities of libEnsemble. libEnsemble has been developed primarily to support research on High-Performance computers, with potentially hundreds of workers performing calculations simultaneously. Please read our platform guides for introductions to using libEnsemble on many such machines.
libEnsemble’s Executors can launch non-Python user applications and simulations across allocated compute resources. Try out this feature with a more-complicated libEnsemble use-case within our Electrostatic Forces tutorial.